vanderPloeg2024_analysis
Source:vignettes/vanderPloeg2024_analysis.Rmd
vanderPloeg2024_analysis.RmdProcessing the data cube
The data cube in vanderPloeg2024$data contains unprocessed counts.
The function processDataCube() performs the processing of
these counts with the following steps:
- It performs feature selection based on the sparsityThreshold
setting. Sparsity is here defined as the fraction of samples where a
microbial abundance (ASV/OTU or otherwise) is zero. For
vanderPloeg2024we can take the RFgroups groups into account for feature selection. We do this by calculating the sparsity for each feature in each subject group and compare those against the sparsity threshold that we set. If a feature passes the threshold in either group, it is selected. - It performs a centered log-ratio transformation of each sample using
the
compositions::clr()function with a pseudo-count of one (on all features, prior to selection based on sparsity). - It centers and scales the three-way array. This is a complex topic that is elaborated upon in our accompanying paper. By centering across the subject mode, we make the subjects comparable to each other within each time point. Scaling within the feature mode avoids the PARAFAC model focusing on features with abnormally high variation.
The outcome of processing is a new version of the dataset. Please
refer to the documentation of processDataCube() for more
information.
processedPloeg = processDataCube(vanderPloeg2024, sparsityThreshold=0.50, considerGroups=TRUE, groupVariable="RFgroup", CLR=TRUE, centerMode=1, scaleMode=2)Determining the correct number of components
A critical aspect of PARAFAC modelling is to determine the correct
number of components. We have developed the functions
assessModelQuality() and
assessModelStability() for this purpose. First, we will
assess the model quality and specify the minimum and maximum number of
components to investigate and the number of randomly initialized models
to try for each number of components.
Note: this vignette reflects a minimum working example for analyzing
this dataset due to computational limitations in automatic vignette
rendering. Hence, we only look at 1-3 components with 5 random
initializations each. These settings are not ideal for real datasets.
Please refer to the documentation of assessModelQuality()
for more information.
# Setup
# For computational purposes we deviate from the default settings
minNumComponents = 1
maxNumComponents = 3
numRepetitions = 5 # number of randomly initialized models
numFolds = 5 # number of jack-knifed models
ctol = 1e-6
maxit = 250
numCores = 1
colourCols = c("RFgroup", "Phylum", "")
legendTitles = c("RF group", "Phylum", "")
xLabels = c("Subject index", "Feature index", "Time index")
legendColNums = c(3,5,0)
arrangeModes = c(TRUE, TRUE, FALSE)
continuousModes = c(FALSE,FALSE,TRUE)
# Assess the metrics to determine the correct number of components
qualityAssessment = assessModelQuality(processedPloeg$data, minNumComponents, maxNumComponents, numRepetitions, ctol=ctol, maxit=maxit, numCores=numCores)We will now inspect the output plots of interest for
vanderPloeg2024.
qualityAssessment$plots$overview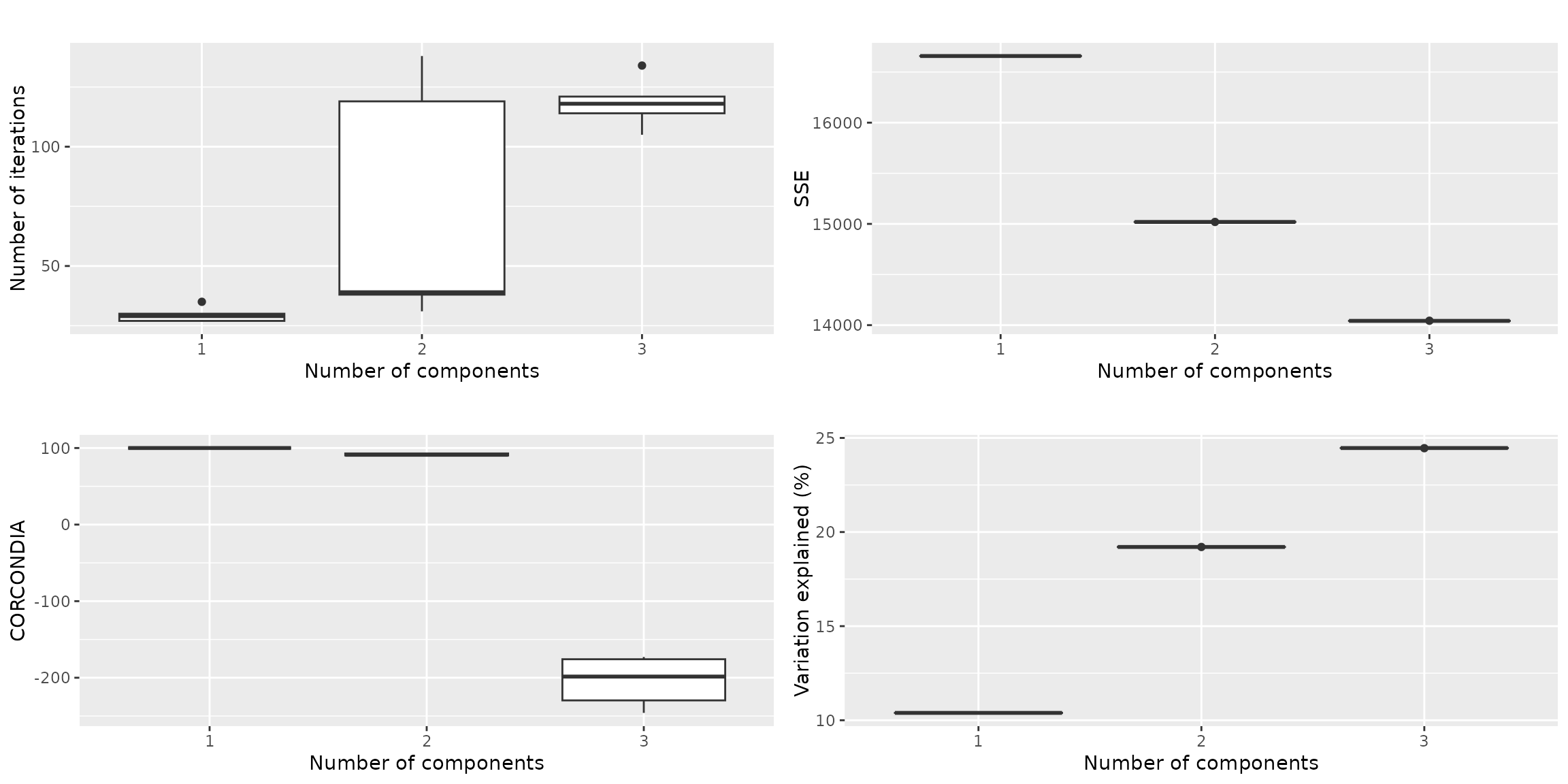 This seems a clear-cut case for a two-component model, as the
three-component models have a CORCONDIA of -100. The maximum amount of
variation we can describe is ~20%.
This seems a clear-cut case for a two-component model, as the
three-component models have a CORCONDIA of -100. The maximum amount of
variation we can describe is ~20%.
Jack-knifed models
Next, we investigate the stability of the models when jack-knifing
out samples using assessModelStability(). We briefly
inspect the two-component and three-component cases although we already
know which will be more appropriate for this data.
stabilityAssessment = assessModelStability(processedPloeg, minNumComponents=1, maxNumComponents=3, numFolds=numFolds, considerGroups=TRUE,
groupVariable="Delivery_mode", colourCols, legendTitles, xLabels, legendColNums, arrangeModes,
ctol=ctol, maxit=maxit, numCores=numCores)
stabilityAssessment$modelPlots[[1]]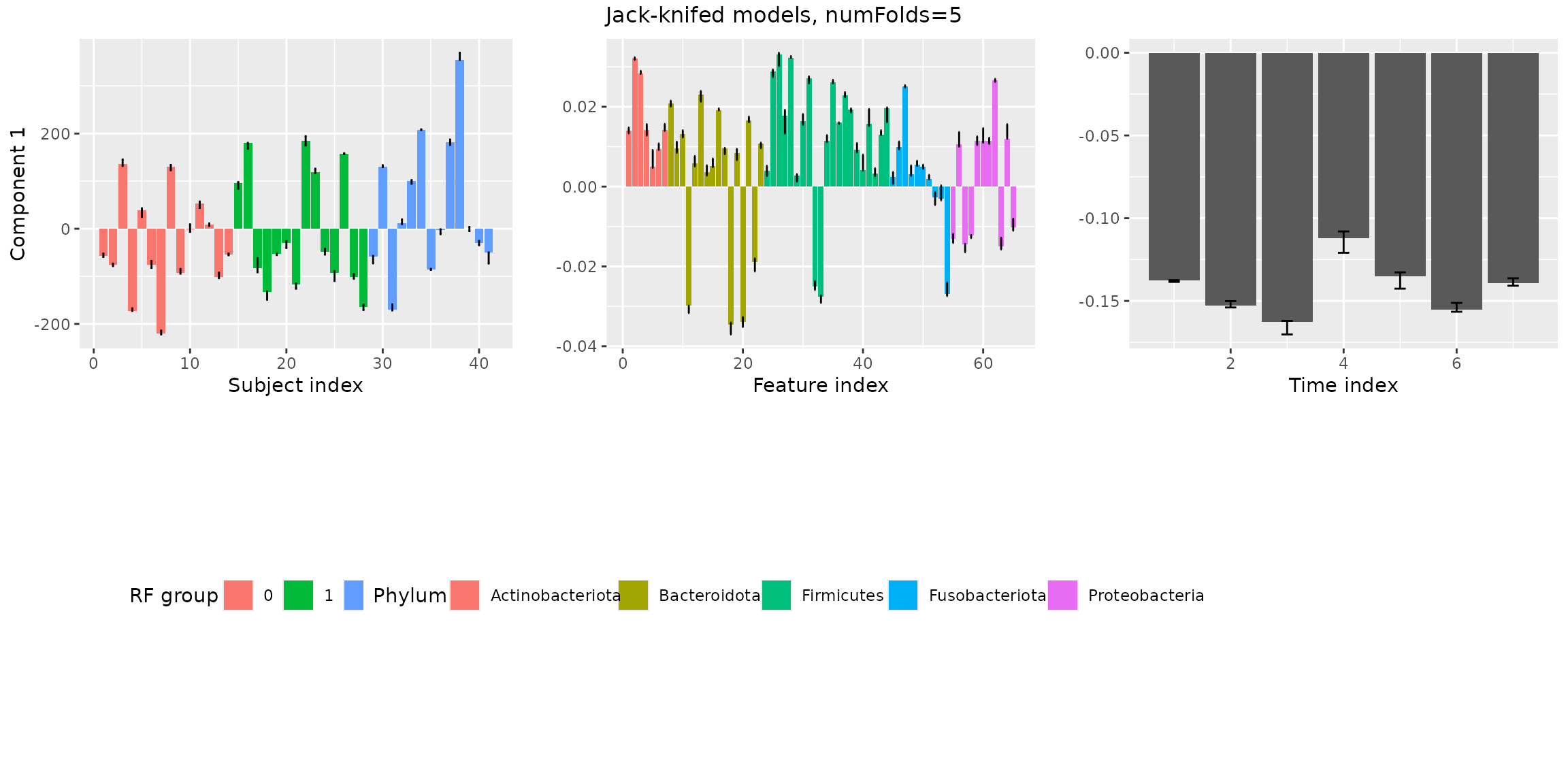
stabilityAssessment$modelPlots[[2]]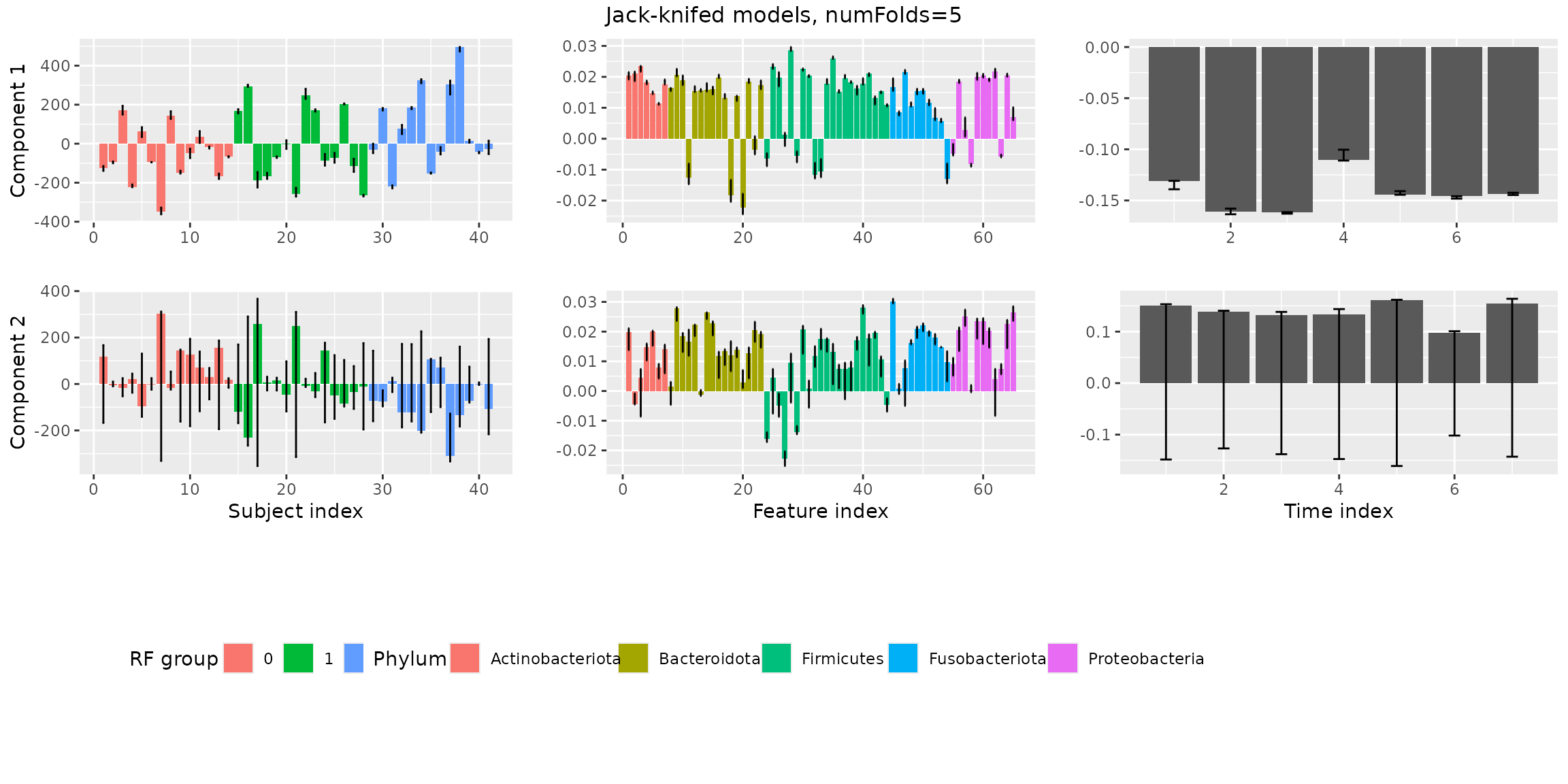
stabilityAssessment$modelPlots[[3]]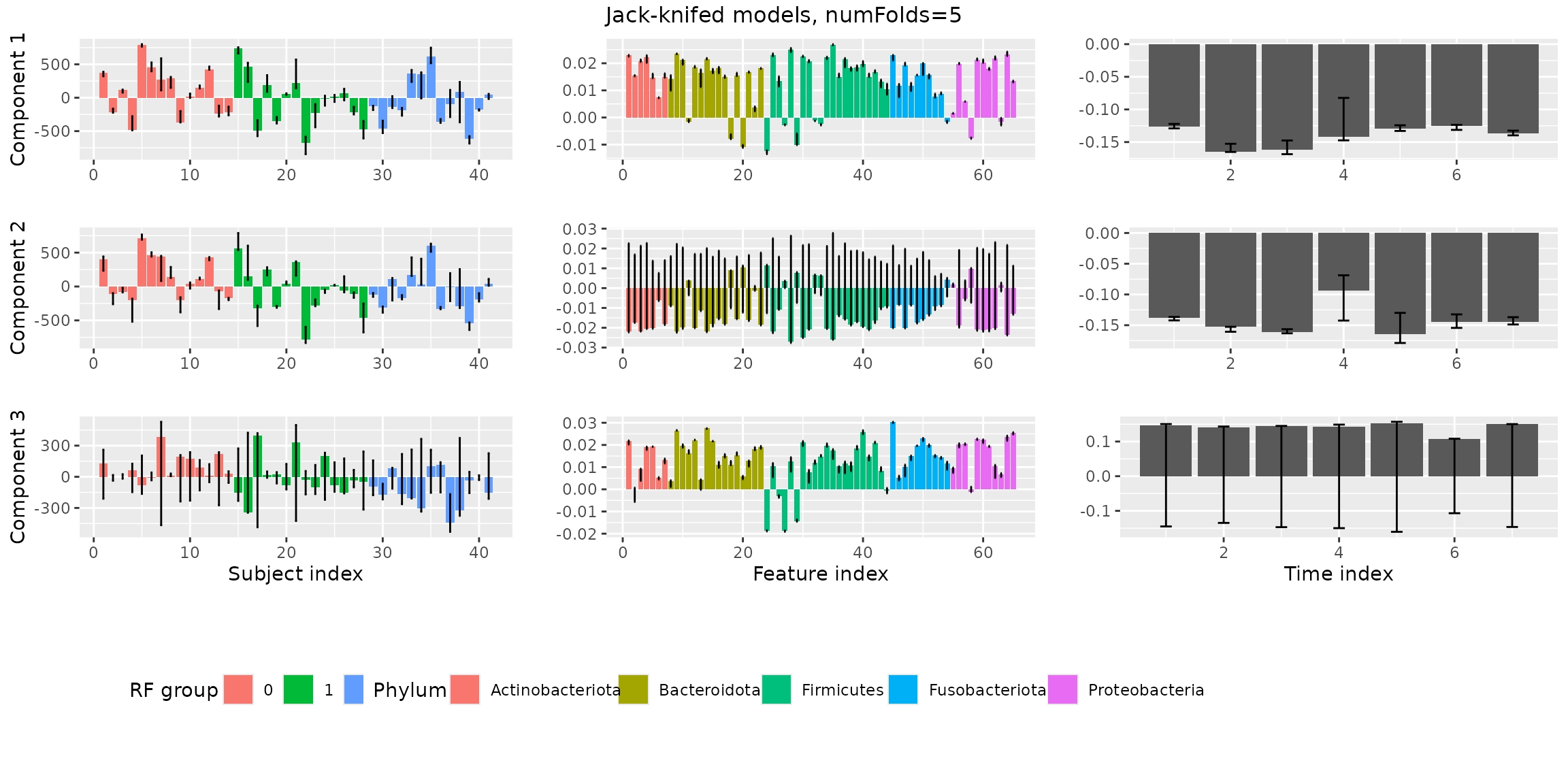
The two-component models are very stable. The three-component models are clearly showing some instability.
Model selection
We have decided that a two-component model is the most appropriate
for the vanderPloeg2024 dataset. We can now select one of
the random initializations from the assessModelQuality()
output as our final model. We’re going to select the random
initialization that corresponded the maximum amount of variation
explained for two components.
numComponents = 2
modelChoice = which(qualityAssessment$metrics$varExp[,numComponents] == max(qualityAssessment$metrics$varExp[,numComponents]))
finalModel = qualityAssessment$models[[numComponents]][[modelChoice]]Finally, we visualize the model using
plotPARAFACmodel().
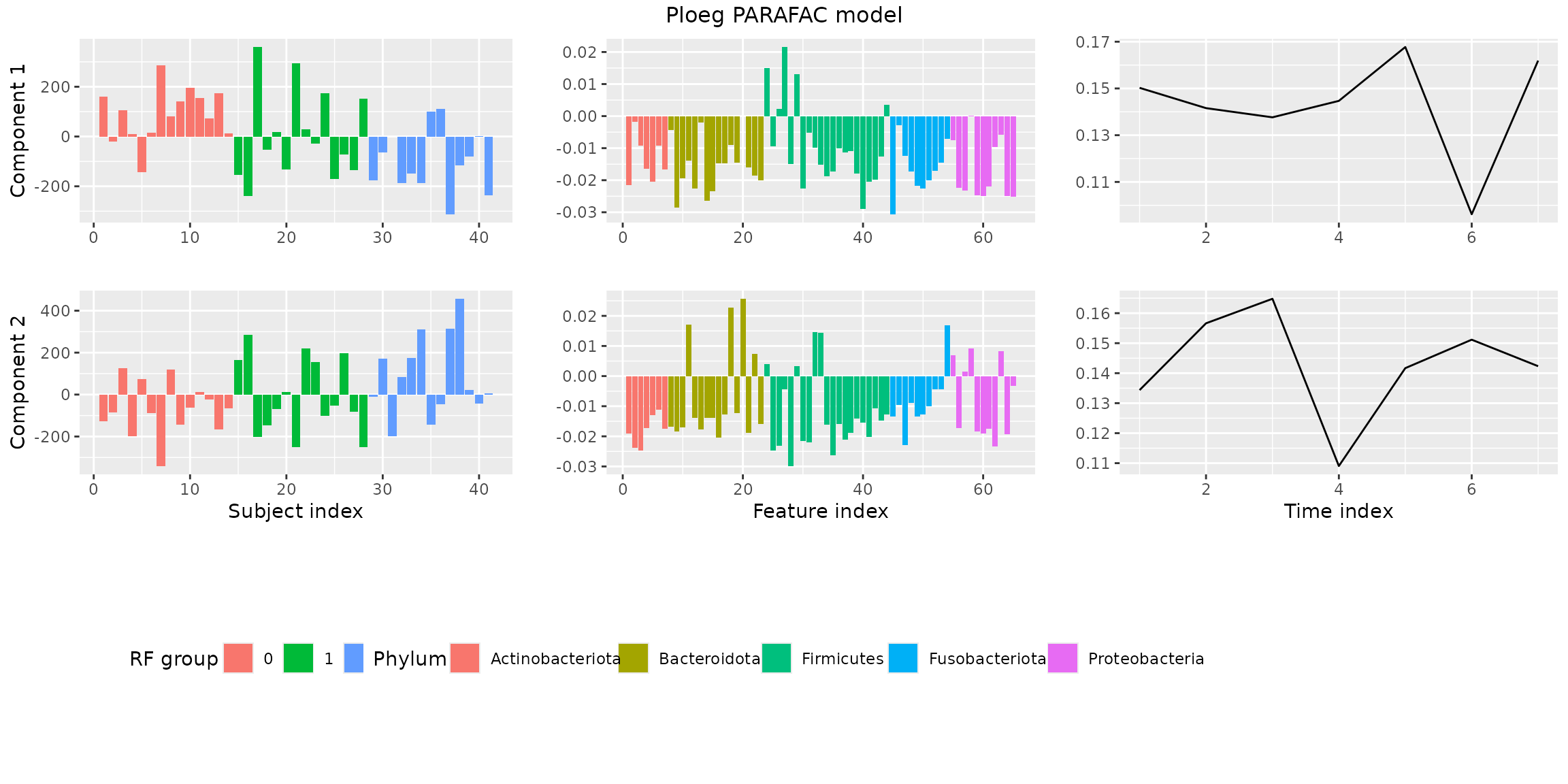
plotPARAFACmodel(finalModel$Fac, processedPloeg, 2, colourCols, legendTitles, xLabels, legendColNums, arrangeModes, continuousModes,
overallTitle = "vanderPloeg2024 PARAFAC model")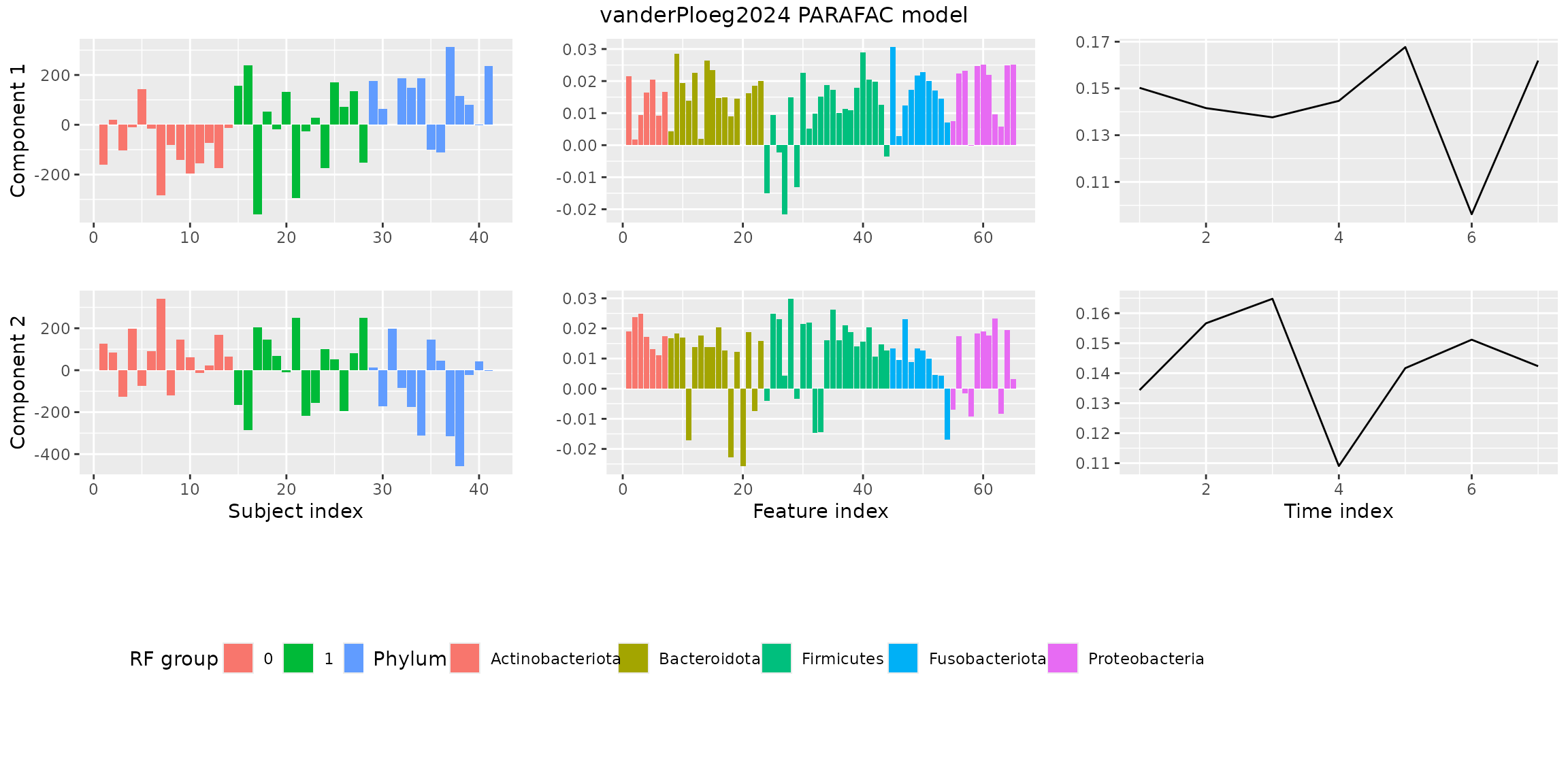
You will observe that the loadings for some modes in some components are negative. This is due to sign flipping: two modes having negative loadings cancel out but describe the same subspace as two positive loadings. We can manually sign flip these loadings to obtain a more interpretable plot.
finalModel$Fac[[1]] = -1 * finalModel$Fac[[1]] # all of mode 1
finalModel$Fac[[2]] = -1 * finalModel$Fac[[2]] # all of mode 2
plotPARAFACmodel(finalModel$Fac, processedPloeg, 2, colourCols, legendTitles, xLabels, legendColNums, arrangeModes,
continuousModes = c(FALSE,FALSE,TRUE),
overallTitle = "Ploeg PARAFAC model")